Replication: The remarkable gymnastics of ORC
Eukaryotic cells that are getting ready to multiply must first faithfully replicate the long DNA molecules that form their genome. This duplication process requires a carefully choregraphed ballet of molecular actors intervening at precisely the right time and place.
For example, in the yeast Saccharomyces cerevisiae – the best characterized DNA replication system to date in eukaryotes (Attali et al., 2021) – the first step consists of the Origin Recognition Complex (or ORC) attaching to specific ‘replication origin’ sequences in the genome (Bell and Stillman, 1992). This process requires binding of a tiny molecule known as ATP. Once in place, ORC can recruit other proteins to build a pre-replicative complex, which cells can activate when they are ready to duplicate their DNA.
The formation of this pre-replicative complex begins with ORC recruiting Cdc6, an enzyme that can also bind ATP. This ORC-Cdc6 structure loads two ring-shaped, hexamer complexes known as Mcm2-7 on to the DNA. Later, these Mcm2-7 hexamers form the functional core of two separate DNA helicases; these enzymes move in opposing directions from the replication origin to unwind the DNA helix, making it accessible to the DNA synthesis machinery (Attali et al., 2021; Noguchi et al., 2017). A gap in the barrel-shaped Mcm2-7 hexamers is kept open by the Cdt1 chaperone so that, when ORC-Cdc6 recruits the Mcm2-7-Cdt1 complex, it can feed double stranded DNA into this opening (Zhai et al., 2017; Yuan et al., 2020). The first Mcm2-7 hexamer then has double stranded DNA passing through its central channel (Yuan et al., 2017). While the identity of the components forming the pre-replicative complex are well known – ORC, Cdc6, Mcm2-7, Cdt1 – precisely how the second Mcm2-7 hexamer is loaded onto the origin DNA needed clarification.
For instance, it has been long known that ORC attaches to the replication origin in an oriented manner by recognizing a particular ‘A element’ motif; however, it can also bind, in the opposite orientation, to a ‘B2 element’ also present in the origin (Figure 1; Bell and Stillman, 1992). In fact, the assembly of the pre-replicative complex and the subsequent bidirectional DNA replication depend on these inverted ORC binding sites (Coster and Diffley, 2017). A logical conclusion was that two ORC molecules would load the two Mcm2-7 helicase hexamers in opposite directions, so that the two hexamers end up in their required head-to-head configuration. Yet, other data suggest that a single ORC could actually load both Mcm2-7 hexamers.
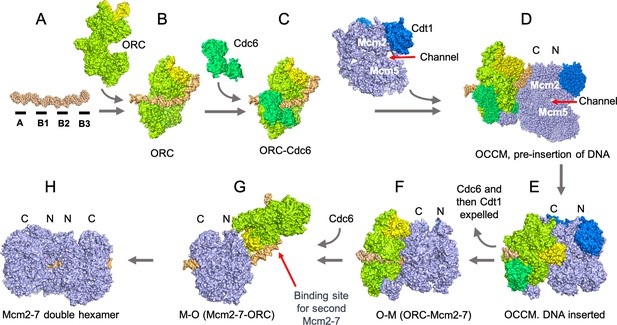
The assembly of the pre-replicative complex requires a seven-step process which involves ORC ‘flipping’ over the Mcm2-7 complex.
(A) The genetic sequence which acts as the starting point for replication in S. cerevisiae (DNA segment in orange) consists of four elements (black segments), with the A and B2 elements binding ORC in opposite orientations. (B) ORC (lime green, Orc6 yellow) first binds to the A and B1 elements. (C) ORC recruits Cdc6 (green). (D) ORC-Cdc6 recruits the Cdt1-Mcm2-7 complex (Cdt1, blue and Mcm2-7 light blue; Mcm2 and Mcm5 are parts of the six proteins in the complex) so that the DNA is aligned to a channel in the Mcm2-7 hexamer. The Mcm2-7 complex is oriented based on the structure of the protein components; with a C-terminus (or extremity; Mcm-C or ‘C’) at one end, and an N-terminus (Mcm-N or ‘N’) at the other. At this stage, Mcm-C binds ORC-Cdc6. (E) The double stranded DNA is inserted into the channel between the Mcm2 and Mcm5 subunits in the Mcm2-7 hexamer and the hexamer is partially closed, to create an intermediary known as the OCCM. (F) ATP hydrolysis by the Mcm2-7 expels the first Cdc6 and then Cdt1 to create the OM complex. (G) ORC flips to the other side of Mcm2-7 and presumably binds to the B2 element, creating the MO complex. As a result, the ORC subunit Orc6 is now orientated towards Mcm2-7-N. (H) ORC can now recruit a second Cdc6, creating a binding site for a second Cdt1-Mcm2-7 complex that is loaded in an opposite orientation to the first Mcm2-7. The Mcm2-7 double hexamer, possibly with ORC still bound to the DNA, establishes the pre-replicative complex that is a precursor for the activation of two enzymes that will unwind the DNA helix when the cell is ready to divide.
Image credit: Molecular structures taken and adapted from the RCSB protein database, deposits 5BK4, 5V8F, 5XF8, 6RQC and 6WGG (Noguchi et al., 2017; Miller et al., 2019; Yuan et al., 2020; Yuan et al., 2017; Zhai et al., 2017).
Examining ‘intermediary’ structures that form as the pre-replicative complex assembles can help to shed light into the mechanisms involved. Such investigations have become possible since scientists have been able to assemble pre-replicative complexes in the laboratory using purified proteins, allowing single-molecule microscopy approaches such as cryo-EM to be paired with genetic and biochemical manipulations (Evrin et al., 2009; Remus et al., 2009; Frigola et al., 2013; Miller et al., 2019; Noguchi et al., 2017; Yuan et al., 2017). As a result, studies have revealed the existence of an intermediate structure known as the OCCM (formed of ORC-Cdc6-Cdt1-Mcm2-7), in which double stranded DNA passes through the middle of the ORC-Cdc6 and Cdt1-Mcm2-7 ring-shaped complexes (Yuan et al., 2017). Other work has highlighted additional intermediate structures consisting of the Mcm2-7-ORC complex bound to DNA in either a MO (Mcm2-7- ORC) or OM (ORC- Mcm2-7) orientation (Miller et al., 2019; Figure 1).
In this study, counting of individual molecules revealed that 74% of the complexes had one ORC, either at the A element or the B2 element, whereas 26% had two. This suggested that loading of the Mcm2-7 double hexamer could occur with either one or two ORCs. Now, in eLife, Stephen Bell and first author Shalini Gupta from the Massachusetts Institute of Technology – with colleagues Jeff Gelles and Larry Friedman at Brandeis University – report new insights that address how one ORC is able to load a second Mcm2-7 hexamer (Gupta et al., 2021).
To conduct their experiments, the team enlisted a replication origin which contained a double stranded DNA segment tethered in a chamber. They labelled various segments of ORC, Cdt1 and Mcm2-7 with different fluorescent compounds; a special type of microscopy called TIRF was then used to track these tagged proteins and how they interact as the pre-replicative complex assembles. The data clearly showed that a single ORC molecule can load two Mcm2-7 hexamers in a head-to-head manner.
To do so, ORC first recruits a Cdc6 protein, and together they load a Cdt1-Mcm2-7 complex – forming the previously identified OCCM complex. ATP is then broken down to release energy, which expels Cdt1 and Cdc6, leaving the MO intermediate bound to the origin. ORC then adds a second Cdc6 which loads another Cdt1-Mcm2-7 hexamer in the opposite direction on the DNA, creating the double Mcm2-7 hexamers in their head-to-head configuration (Figure 1).
The remarkable observation is that ORC switches from one side to the other of the first Mcm2-7 hexamer, presumably jumping between binding to the A and B2 elements. The ‘flip’ creates a new intermediate structure consistent with the MO complex observed previously (Miller et al., 2019; Figure 1). ORC’s impressive gymnastic move is possible because the first Mcm2-7 hexamer breaks down ATP, providing the required energy for the complex to slide along the DNA strand, exposing the B2 element. This allows ORC to then flip orientation, and to load the second Mcm2-7 hexamer in the right position. It remains to be determined how this occurs in other eukaryotic cells, most of which do not have specific genetic sequences similar to the A or B2 elements in their replication origins.
Large ATP-driven molecular movements have been known for some time – a classic example being the motor proteins kinesin or dynein, which can use ATP to change conformation and shuttle molecules by ‘walking’ on the cell’s internal skeleton (Gennerich and Vale, 2009). The results by Gupta et al. provide another, remarkably elegant example of the diverse and intricate ATP-powered feats which keep cells ticking.
References
-
Structural mechanisms for replicating DNA in eukaryotesAnnual Review of Biochemistry 90:77–106.https://doi.org/10.1146/annurev-biochem-090120-125407
-
Walking the walk: how kinesin and dynein coordinate their stepsCurrent Opinion in Cell Biology 21:59–67.https://doi.org/10.1016/j.ceb.2008.12.002
-
Structural basis of Mcm2-7 replicative helicase loading by ORC-Cdc6 and Cdt1Nature Structural & Molecular Biology 24:316–324.https://doi.org/10.1038/nsmb.3372
-
Open-ringed structure of the Cdt1-Mcm2-7 complex as a precursor of the MCM double hexamerNature Structural & Molecular Biology 24:300–308.https://doi.org/10.1038/nsmb.3374
Article and author information
Author details
Acknowledgements
Some of the research cited in this article and used to generate the figure was supported by NIH grants (R01-GM45436 and P01-CA13106).
Publication history
- Version of Record published: February 9, 2022 (version 1)
Copyright
© 2022, Stillman
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
-
- 983
- views
-
- 167
- downloads
-
- 10
- citations
Views, downloads and citations are aggregated across all versions of this paper published by eLife.
Download links
Downloads (link to download the article as PDF)
Open citations (links to open the citations from this article in various online reference manager services)
Cite this article (links to download the citations from this article in formats compatible with various reference manager tools)
Further reading
-
- Biochemistry and Chemical Biology
Transporter research primarily relies on the canonical substrates of well-established transporters. This approach has limitations when studying transporters for the low-abundant micromolecules, such as micronutrients, and may not reveal physiological functions of the transporters. While d-serine, a trace enantiomer of serine in the circulation, was discovered as an emerging biomarker of kidney function, its transport mechanisms in the periphery remain unknown. Here, using a multi-hierarchical approach from body fluids to molecules, combining multi-omics, cell-free synthetic biochemistry, and ex vivo transport analyses, we have identified two types of renal d-serine transport systems. We revealed that the small amino acid transporter ASCT2 serves as a d-serine transporter previously uncharacterized in the kidney and discovered d-serine as a non-canonical substrate of the sodium-coupled monocarboxylate transporters (SMCTs). These two systems are physiologically complementary, but ASCT2 dominates the role in the pathological condition. Our findings not only shed light on renal d-serine transport, but also clarify the importance of non-canonical substrate transport. This study provides a framework for investigating multiple transport systems of various trace micromolecules under physiological conditions and in multifactorial diseases.
-
- Biochemistry and Chemical Biology
- Cell Biology
Mediator of ERBB2-driven Cell Motility 1 (MEMO1) is an evolutionary conserved protein implicated in many biological processes; however, its primary molecular function remains unknown. Importantly, MEMO1 is overexpressed in many types of cancer and was shown to modulate breast cancer metastasis through altered cell motility. To better understand the function of MEMO1 in cancer cells, we analyzed genetic interactions of MEMO1 using gene essentiality data from 1028 cancer cell lines and found multiple iron-related genes exhibiting genetic relationships with MEMO1. We experimentally confirmed several interactions between MEMO1 and iron-related proteins in living cells, most notably, transferrin receptor 2 (TFR2), mitoferrin-2 (SLC25A28), and the global iron response regulator IRP1 (ACO1). These interactions indicate that cells with high MEMO1 expression levels are hypersensitive to the disruptions in iron distribution. Our data also indicate that MEMO1 is involved in ferroptosis and is linked to iron supply to mitochondria. We have found that purified MEMO1 binds iron with high affinity under redox conditions mimicking intracellular environment and solved MEMO1 structures in complex with iron and copper. Our work reveals that the iron coordination mode in MEMO1 is very similar to that of iron-containing extradiol dioxygenases, which also display a similar structural fold. We conclude that MEMO1 is an iron-binding protein that modulates iron homeostasis in cancer cells.






