Hippocampome.org 2.0 is a knowledge base enabling data-driven spiking neural network simulations of rodent hippocampal circuits
Figures
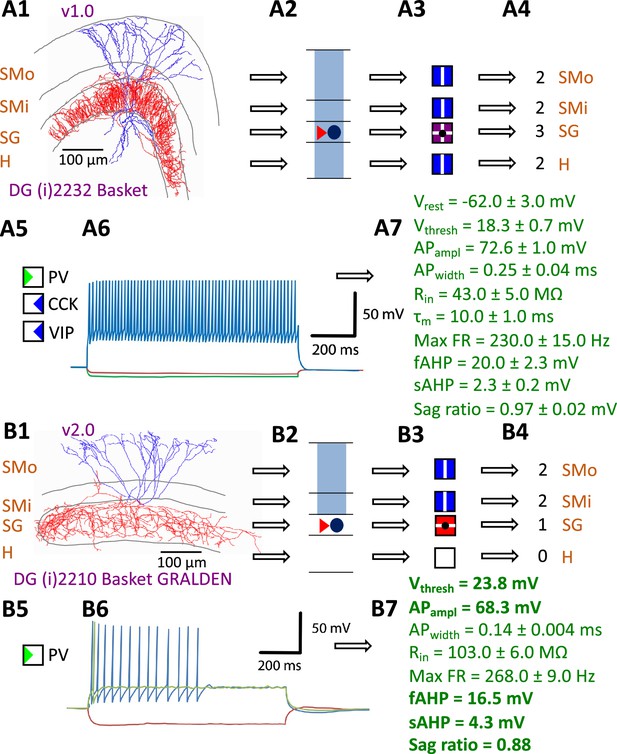
Defining neuron types in Hippocampome.org.
(A) Properties of a Hippocampome.org v1.0 neuron type. (A1) Morphology of a Dentate Gyrus (i) 2232 Basket cell (NeuroMorpho.Org cell NMO_34300: Figure S3A in Hosp et al., 2014) with axons (red) in stratum granulosum (SG) and dendrites (blue) in all four layers. (A2) Schematic interpretation of the morphological tracing, where the circle represents the location of the soma in SG, the red triangle the location of the axons in SG, and the blue rectangles the locations of the dendrites in all four layers. (A3) Hippocampome.org representation of the morphology, where a blue square with a vertical line (|) indicates dendritic presence in the outer two-thirds of the stratum moleculare (SMo) and the inner one-third of the stratum moleculare (SMi) and the hilus (H), a purple square with a cross (+) indicates both axonal and dendritic presence in SG, and a black dot (•) indicates the soma location in SG. (A4) Hippocampome.org numerical coding of the reconstructed neuron, where 2 indicates the presence of dendrites (in SMo, SMi, and H); and 3 indicates the presence of both axons and dendrites (in SG). (A5) Biomarker expressions, where a green triangle indicates positive expression for parvalbumin (PV), and blue triangles indicate negative expression for cholecystokinin (CCK) and vasoactive intestinal polypeptide (VIP). (A6) Firing pattern phenotype (non-adapting spiking (NASP); adapted from Figure 1B1 in Savanthrapadian et al., 2014). (A7) Membrane biophysics values (from Figure 3C and Table 1 in Lübke et al., 1998) recorded at 35–37°C. (B) Properties for a Hippocampome.org v2.0 neuron type. (B1) Morphology of a DG (i) 2210 Basket GRALDEN (NeuroMorpho.Org cell NMO_146159: Figure 4S3 in Vaden et al., 2020) with red axons in SG and blue dendrites in SMo and Smi. (B2) Schematic interpretation of the reconstruction (same symbols as in A2). (B3-4) Hippocampome.org representation and numerical coding of the morphology (same symbols as in A3-4). (B5) Biomarker expression. (B6) Firing pattern phenotype (silence preceded by transient stuttering (TSTUT.SLN); adapted from Figure S4 in Markwardt et al., 2011). (B7) Membrane biophysics values recorded at room temperature (from Figure 4D in Vaden et al., 2020), and at 22 °C (from Figure S4 in Markwardt et al., 2011); emboldened values were extracted from the firing pattern trace in B6. Membrane biophysics abbreviations: Vrest: resting membrane potential; Vthresh: firing threshold potential; APampl: action potential amplitude; APwidth: action potential width; Rin: input resistance; τm: membrane time constant; Max FR: maximum firing rate; fAHP: fast after-hyperpolarizing potential; sAHP: slow after-hyperpolarizing potential; Sag ratio: ratio of the steady-state membrane potential to the minimum membrane potential.
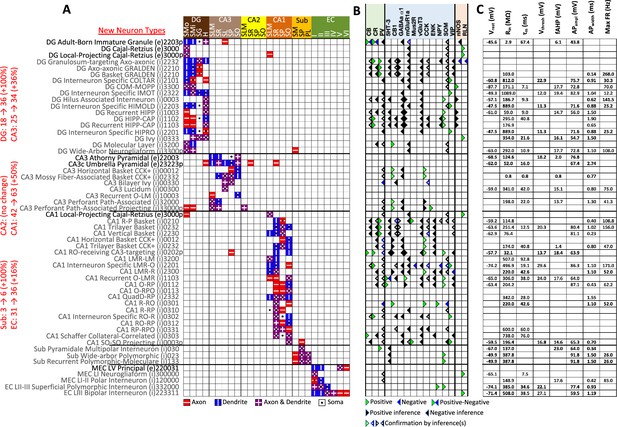
New neuron types added to Hippocampome.org v2.0.
(A) Morphology encodings of the 56 new neuron types that extend the 124 types in Hippocampome.org v1.12. (Left) Increase in number of neuron types for each subregion. For the neuron type names, excitatory types (e) are in black font and inhibitory types (i) are in gray font. The 3–6 digit numbers encode the patterns of axons and dendrites in the layers of the home subregion of the neuron type: 0=no axons or dendrites, 1=only axons, 2=only dendrites, 3=both axons and dendrites. A “p” indicates that the neuron types projects across into other subregions. (B) Biomarker expressions of the neuron types. (C) Membrane biophysics values for the neuron types. Morphology abbreviations: DG: dentate gyrus; SMo: outer two-thirds of stratum moleculare; SMi: inner one-third of stratum moleculare; SG: stratum granulosum; H: hilus; SLM: stratum lacunosum-moleculare; SR: stratum radiatum; SL: stratum lucidum; SP: stratum pyramidale; SO: stratum oriens; Sub: subiculum; PL: polymorphic layer; EC: entorhinal cortex. Marker abbreviations: CB: calbindin; CR: calretinin; PV: parvalbumin; 5HT-3: serotonin receptor 3; CB1: cannabinoid receptor type 1; GABAa α1: GABA-a alpha 1 subunit; mGluR1a: metabotropic glutamate receptor 1 alpha; Mus2R: muscarinic type 2 receptor; vGluT3: vesicular glutamate transporter 3; CCK: cholecystokinin; ENK: enkephalin; NPY: neuropeptide Y; SOM: somatostatin; VIP: vasoactive intestinal polypeptide; nNOS: neuronal nitric oxide synthase; RLN: reelin. Membrane biophysics abbreviations: see Figure 1.
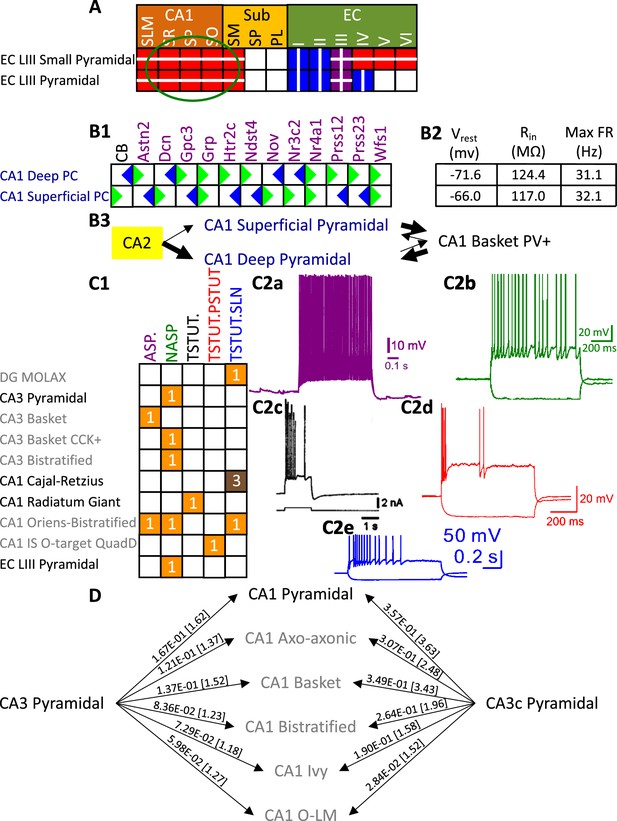
Extensions to the neuronal properties of Hippocampome.org v1.x.
(A) Additions to the axonal projections (circled in green) for two v1.0 neuron types, based on information derived from Figure 2b in Deller et al., 1996. (B1) Biomarker expressions for the two CA1 Pyramidal sub-types added in Hippocampome.org v1.9 (Sanchez-Aguilera et al., 2021). (B2) Membrane biophysics values for the two sub-types. (B3) CA2 projects preferentially to the deep sublayer of CA1 (Kohara et al., 2014). More perisomatic parvalbumin-positive (PV+) GABAergic boutons are found at CA1 Deep Pyramidal cells (Valero et al., 2015). CA1 Superficial Pyramidal cells form more frequent connections to PV + CA1 Basket cells, and PV + CA1 Basket cells form significantly more perisomatic axon terminals on CA1 Deep Pyramidal cells (Lee et al., 2014). (C1) Additions to the firing pattern phenotypes of v1.0 neuron types. (C2a) Example of adapting spiking (ASP.) in a CA1 Oriens-Bistratified cell (adapted from Figure 4B in Craig and McBain, 2015). (C2b) Example of non-adapting spiking (NASP) in a CA3 Basket CCK + cell (adapted from Figure 3A in Szabadics and Soltesz, 2009). (C2c) Example of transient stuttering (TSTUT.) in a CA1 Radiatum Giant cell (reproduced from Figure 2Bb in Kirson and Yaari, 2000). (C2d) Example of transient stuttering followed by persistent stuttering (TSTUT.PSTUT) in a CA1 Interneuron Specific O-targeting QuadD (adapted from Figure 2D in Chamberland et al., 2010). (C2e) Example of silence preceded by transient stuttering (TSTUT.SLN) in a DG MOLAX cell (adapted from Figure S2c in Lee et al., 2016). (D) Synaptic probabilities for projecting connections and the corresponding number of contacts in brackets between the two CA3 Pyramidal neuron types and a selection of CA1 neuron types. Morphology abbreviations: see Figure 2. Marker abbreviations: CB: calbindin; Astn2: astrotactin 2; Dcn: decorin; Gpc3: glypican 3; Grp: gastrin releasing peptide; Htr2c: 5-hydroxytryptamine receptor 2 c; Ndst4: N-deacetylase and N-sulfotransferase 4; Nov: nephroblastoma overexpressed; Nr3c2: nuclear receptor subfamily 3 group C member 2; Nr4a1: nuclear receptor subfamily 4 group A member 1; Prss12: serine protease 12; Prss23: serine protease 23; Wfs1: wolframin ER transmembrane glycoprotein. Membrane biophysics abbreviations: see Figure 1.
© 2000, Society for Neuroscience. Figure 3C2c is reproduced from Figure 2Bb from Kirson and Yaari, 2000, with permission from Society for Neuroscience. It is not covered by the CC-BY 4.0 licence and further reproduction of this panel would need permission from the copyright holder.
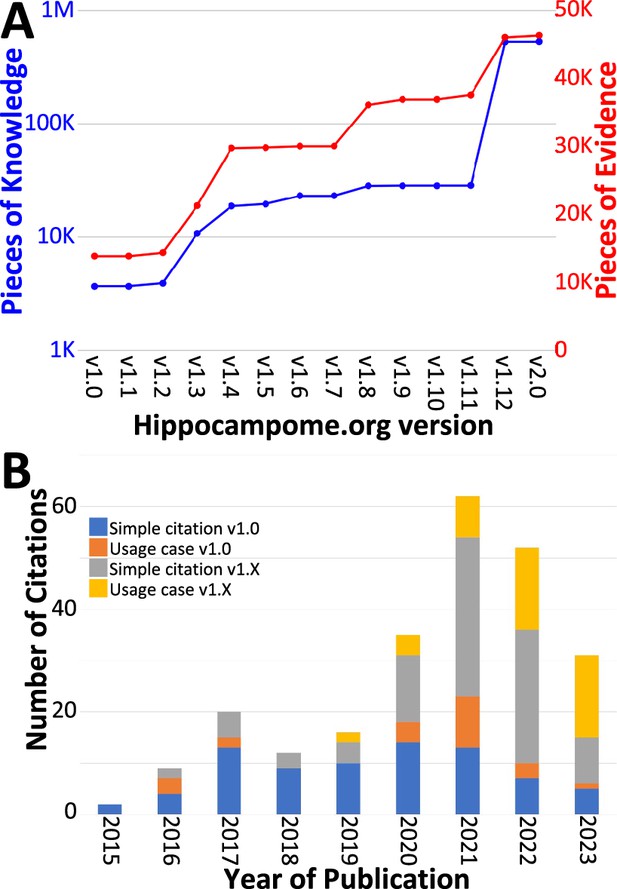
Trends in Hippocampome.org data, knowledge, citations, and usage since v1.0.
(A) Increase in pieces of knowledge (blue) and evidence (red) with Hippocampome.org version number. (B) Number of citations, in which the publication is simply referenced (blue and gray portions), and usage cases, in which the citing work makes use of the information contained within the Hippocampome.org-related work (orange and yellow portions), by year.
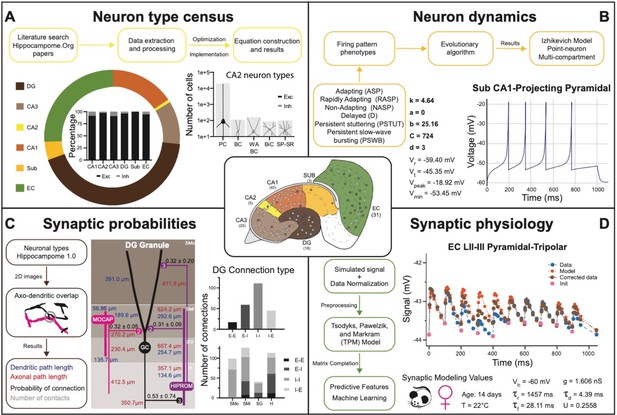
Transitional knowledge enabling Hippocampome.org to support spiking neural network simulations.
Center: General diagram of the hippocampal formation and the number of cell types in Hippocampome.org v1.0. (A) Neuron type census. Top: General pipeline for obtaining cell counts for specific collections of neurons from the peer reviewed literature. Left: Neuron count proportions for the different subregions of the hippocampal formation. Insert: Normalized neuron counts for the inhibitory vs. excitatory balance by subregion. Right: Neuron counts for five identified CA2 neuron types (green schematic: excitatory, red schematic: inhibitory). (B) Neuron dynamics. Top: General pipeline for obtaining Izhikevich models to reproduce the firing pattern phenotypes from peer reviewed data. Right: Simulated firing pattern from a Sub CA1-Projecting Pyramidal cell in response to a 250 pA current injection pulse lasting 1 s. Izhikevich model parameters are shown in bold and the membrane biophysics properties are shown in regular font. (C) Synaptic probabilities. Left: General pipeline for obtaining the connection probabilities, number of contacts, and dendritic and axonal path lengths from 2D reconstructions. Middle: Example of a connectivity diagram of a DG Granule cell and two interneurons across the different parcels of DG. Probabilities of connection (mean ± SD) are shown in black, numbers of contacts in gray, dendritic path lengths in blue, and axonal lengths in red. Right top: Total number of connections within DG by connection type. Right bottom: Breakdown of the total number of connections by parcel and connection type. (D) Synaptic physiology. Left: General pipeline for obtaining normalized synaptic parameters from paired recordings with a TPM model. Right top: Digitized synaptic data between two EC LII-III Pyramidal-Tripolar cells. Experimental data are shown in blue, initiation synaptic points in pink, model data in orange, and corrected data in green. Right bottom: Simulated modeling conditions, electrophysiological parameters, and TPM parameters. Abbreviations by panel: (A) PC: Pyramidal cell; BC: Basket cell; WA BC: Wide-Arbor Basket cell; BiC: Bistratified cell; Exc: excitatory; Inh: inhibitory. (B) Vr: resting membrane potential; Vt: firing threshold potential; Vpeak: spike cutoff potential; Vmin: post-spike reset potential. (C) E-E: excitatory-excitatory; E-I: excitatory-inhibitory; I-I: inhibitory-inhibitory; I-E: inhibitory-excitatory. (D) Vh: holding potential; τr: recovery time constant; τf: facilitation time constant; g: conductance; τd: deactivation time constant; U: utilization ratio.
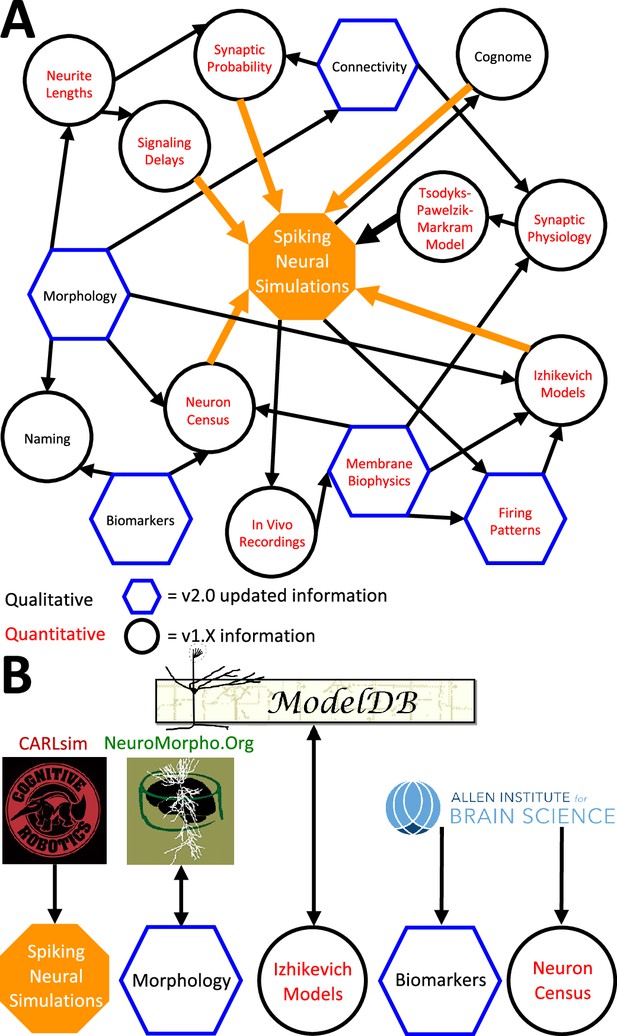
Hippocampome.org data provenance.
(A) The internal web of constituent neuron-type properties (black thin arrows) that ultimately contribute to the instantiation of spiking neural simulations (orange thick arrows). Properties described qualitatively, such as morphological presence of axons in a layer or molecular biomarker expressions, are in black font. Properties described by quantitative values, such as membrane biophysics and neurite lengths, are in red font. Properties with v2.0 updated information, such as connectivity and firing pattern phenotypes, are depicted by blue hexagons, and v1.x information, such as Izhikevich modeling parameter values and neuron-type census values, is visualized by black circles. (B) External resources that contribute data to and receive data from Hippocampome.org (the ModelDB logo has been modified from the original).
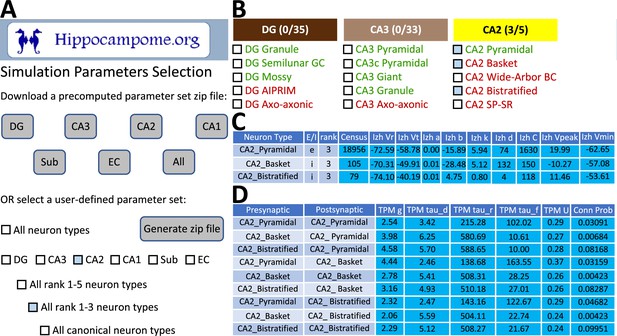
CARLsim simulation parameters selection and file generation interface.
(A) The user chooses which subset of the available neuron types to include in the generated downloadable parameter file. Neuron types can be selected (check boxes and gray highlights) either individually or by groupings, such as by subregion and/or by importance rank. (B) Representative user selection. (C) Downloadable neuron-level parameters. (D) Downloadable connection-level parameters.
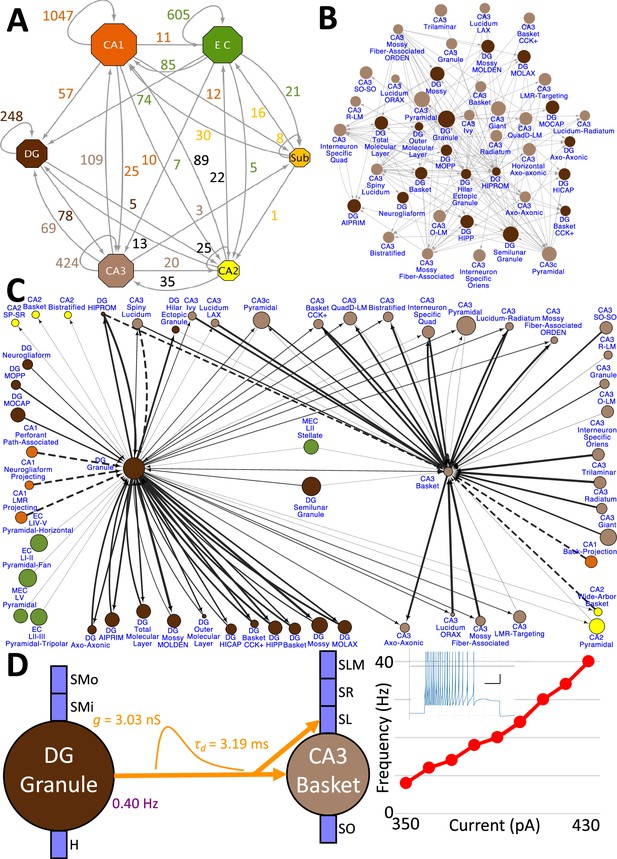
Hierarchy of neuronal connectivity in Hippocampome.org.
(A) Subregional connectivity, where the number of connections between subregions is shown, and the node size is proportional to the number of neuron types in each subregion. (B) The reciprocal connectivity between DG and CA3 neuron types consists of 147 connections. The node size is proportional to the census size for each neuron type. (C) The full connectivity involving DG Granule and CA3 Basket neuron types consists of 98 connections. The node size is proportional to the census size for each neuron type, and the thicknesses of the connecting arrows are proportional to the synaptic probability. The dashed lines are connections for which the synaptic probability has been approximated based on the means of known values. (D) The electrophysiological connection between a DG Granule cell and a CA3 Basket cell. The in vivo firing rate is shown for the presynaptic neuron. The transfer function between the two neuron types is proportional to the synaptic conductance times the single-exponential decay time constant (g · τd; rat, male, P56, 32 °C, current clamp). The frequency-current (F–I) curve of the single-compartment Izhikevich model of a CA3 Basket cell was obtained with 10 pA current steps. Inset: Izhikevich model firing pattern of a CA3 Basket cell simulated with 430 pA of current applied for 500 ms (vertical and horizontal scale bars, respectively).
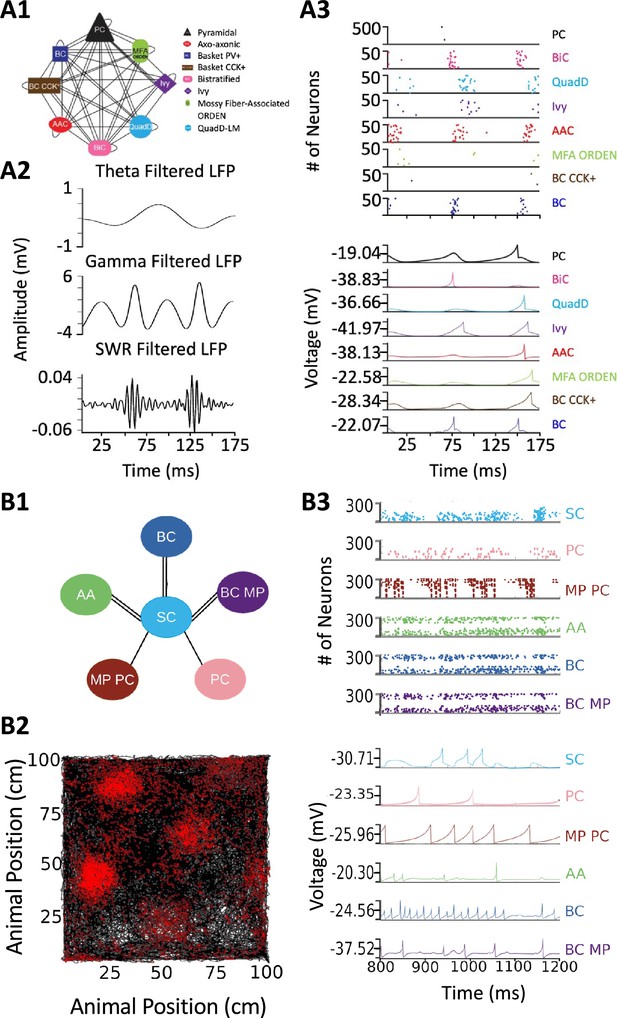
Spiking neural network simulations.
(A) Full-scale CA3 model. (A1) Neuron type connectivity schematic. (A2) Theta (4–12 Hz; top), Gamma (25–100 Hz; middle), and Sharp-Wave Ripple (150–200 Hz; bottom) filtered local field potentials from 175ms of the simulation. (A3) Raster plot of 500 Pyramidal cells and 50 interneurons of each type (top), and representative voltage traces for each neuron type (bottom) during the same 175ms of the simulation in (A2). (B) A mock up of a spatial representation through grid cell firing. (B1) Neuron type connectivity schematic. (B2) Simulated animal trajectory (black) with red dots indicating the firing of a neuron in those locations. (B3) Raster plot of 300 neurons from each type (top), and representative voltage traces for each neuron type. Abbreviations by panel: (A) PC: Pyramidal cell; BiC: Bistratified cell; QuadD: QuadD-LM cell; AAC: Axo-axonic cell; MFA ORDEN: Mossy Fiber-Associated ORDEN cell; BC CCK+: cholecystokinin-positive Basket cell. (B) SC: Medial Entorhinal Cortex Layer II Stellate cell; PC: Entorhinal Cortex Layer III Pyramidal cell; MP PC: Entorhinal Cortex Layer I-II Multipolar Pyramidal cell; AA: Entorhinal Cortex Layer II Axo-axonic; BC: Medial Entorhinal Cortex Layer II Basket; BC MP: Entorhinal Cortex Layer II Basket-Multipolar Interneuron.
Tables
Added knowledge and functioning in Hippocampome.org releases v1.1–12.
| Version | Contribution | Article |
|---|---|---|
| v1.1 |
| Hamilton et al., 2017a |
| v1.2 |
| Rees et al., 2016 |
| v1.3 |
| Hamilton et al., 2017b |
| v1.4 |
| Moradi and Ascoli, 2020 |
| v1.5 |
| White et al., 2020 |
| v1.6 |
| Komendantov et al., 2019 |
| v1.7 |
| Venkadesh et al., 2019 |
| v1.8 |
| Tecuatl et al., 2021b |
| v1.9 |
| Sanchez-Aguilera et al., 2021 |
| v1.10 |
| Sutton and Ascoli, 2021 |
| v1.11 |
| Attili et al., 2022 |
| v1.12 |
| Moradi et al., 2022 |
Probabilities of connection and number of contacts per connected pair from DG Granule cell to mossy fiber targets in CA3.
| Postsynaptic neuron type | Probability | # contacts |
|---|---|---|
| CA3 Pyramidal | 1.11E-04 | 1.08 |
| CA3c Pyramidal | 3.91E-04 | 1.31 |
| CA3 Spiny Lucidum Dentate-Projecting | 5.89E-04 | 1.69 |
| CA3 Mossy Fiber-Associated ORDEN | 4.44E-04 | 1.27 |
| CA3 Basket | 6.55E-04 | 1.50 |
| CA3 Basket CCK+ | 2.14E-04 | 1.16 |
| CA3 Ivy | 3.35E-04 | 1.29 |
| CA3 Mossy Fiber-Associated | 3.78E-05 | 1.04 |
| CA3 LMR-Targeting | 1.31E-04 | 1.21 |
| CA3 Lucidum ORAX | 2.62E-04 | 1.19 |
| CA3 Lucidum-Radiatum | 3.25E-04 | 1.13 |
| CA3 Axo-Axonic | 7.56E-04 | 1.50 |
| CA3 Bistratified | 8.25E-04 | 1.45 |
| CA3 QuadD-LM | 2.91E-04 | 1.25 |
Examples of independent studies utilizing unique neuronal properties from Hippocampome.org v1.0.
| Article | Usage |
|---|---|
| Gulyás et al., 2016 | Lists of subthreshold physiological properties for multicompartmental modeling |
| Skene and Grant, 2016 | Catalog of CA1 Interneuron types |
| Faghihi and Moustafa, 2017 | Diversity of hippocampal neuron types and morphological neuronal features |
| Puighermanal et al., 2017 | Biomarker expression in CA1 interneurons |
| Depannemaecker et al., 2020 | Parameter values for a model of synaptic neurotransmission |
| Ecker et al., 2020 | Evidence that CA1 interneurons express multiple overlapping chemical markers |
| Hunsberger and Mynlieff, 2020 | Cell identification based on firing properties |
| Schumm et al., 2020 | Directionality of connections in the hippocampus |
| Aery Jones et al., 2021 | Local connectivity of CA1 PV + interneurons |
| Ciarpella et al., 2021 | Lists of hippocampal genes |
| Luo et al., 2021 | Confirmation of multiple hippocampal neuron types |
| Mehta et al., 2021 | Connectome model inspired by entorhinal-CA1 circuit |
| Obafemi et al., 2021 | Principal channels of information processing are DG Granule cells and CA1-3 Pyramidal cells |
| Sáray et al., 2021 | Membrane biophysics values for CA1 Pyramidal cells |
| Smith et al., 2021 | Omni-directionality of axons of CA1 Pyramidal cells |
| Venkadesh and Van Horn, 2021 | Example of a brain region’s mesoscopic structural connectivity |
| Walker et al., 2021 | Reference to morphological and molecular characteristics of hippocampal principal cells and interneurons |
| Wynne et al., 2021 | Example brain region with a variety of cell types |
| Kopsick et al., 2023 | Utilize accumulated knowledge as the basis for simulations |
| Schumm et al., 2022 | Hippocampal morphology, biomarker expression, connectivity, and typing of neurons |
| Zagrean et al., 2022 | Diversity of hippocampal neuronal types and their properties |





